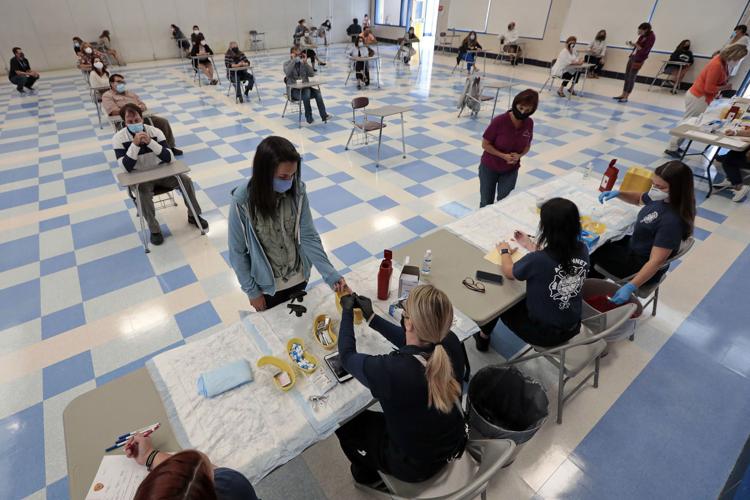
Early detection and intervention efforts were successful at containing the first cases of coronavirus in North America and Europe, but insufficient testing and tracing created new transmission opportunities and ignited a pandemic that has killed 950,000 people worldwide, a new University of Arizona-led study found.
By mapping the genomes of thousands of coronavirus samples and analyzing detailed travel records, researchers used computer simulations to reconstruct the global spread of the virus from its origins in Wuhan, China. The study pinpoints the introduction of the virus into each country and reveals which early public-health measures were effective and which failed to stop the viral spread.
“Our aspiration was to develop and apply powerful new technology to conduct a definitive analysis of how the pandemic unfolded in space and time, across the globe,” said UA researcher Michael Worobey, who led an interdisciplinary team of scientists from 13 research institutions in the U.S., Belgium, Canada and the U.K. “Before, there were lots of possibilities floating around in a mishmash of science, social media and an unprecedented number of preprint publications still awaiting peer review.”
People are also reading…
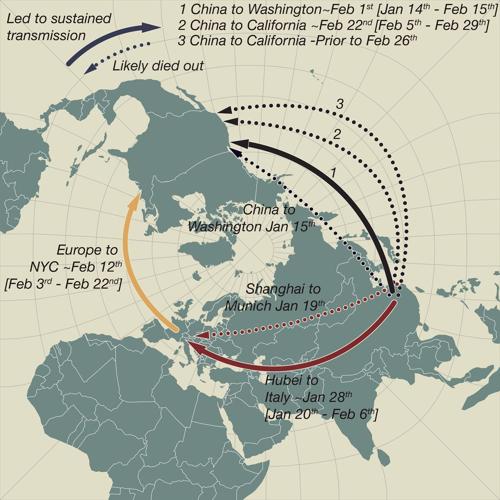
Before the study, most people believed the earliest known cases of COVID-19 — a traveler from Wuhan to Seattle on Jan. 15 and a German businessman returning home from Shanghai on Jan. 20 — marked the beginning of the viral spread in the United States and Europe. However, the research shows both of these patients were effectively isolated and treated, with limited community exposure.
It wasn’t until six weeks later that several additional cases were detected in Washington, which involved viral strains that were genetically different from the two original cases, making it unlikely that the first documented cases led to community transmission.
“All of these things kind of went right with this first patient in Washington. And our study now shows that when you do things right, they can have a very powerful effect of shutting down transmission. And if we had been able to do that with the subsequent cases, we could have prevented the whole outbreak from occurring in the U.S.”
Despite early successes in containment, the findings indicate that additional cases of coronavirus entered the U.S. in early February, carried by travelers from China. The virus went largely undetected for several weeks, resulting in widespread community transmission.
The exact case that led to the larger epidemic is unknown, however. At least 40,000 U.S. citizens and visa holders continued to travel from China to the U.S. even after a travel ban was implemented in early February.
Echoing the early U.S. response, early cases of the virus in Germany were effectively eliminated with proper testing, contact tracing and isolation. According to the study, Germany was not the origin of the epidemic in Europe. Rather, it was China to northern Italy and then to the rest of the continent.
Italy was also the source for the later spread of the virus to New York City and other outbreaks in the U.S., carried again by travelers.
Think of genomes as a necklace
To determine how and when the virus traveled across the world, Worobey and his collaborators used a collection of over 100,000 coronavirus genome samples and identified patterns of mutations.
Genomes are like a necklace with four colors of beads, Worobey said. The genome of coronavirus has 30,000 beads and they have a very specific color arrangement. Those beads get interpreted by our cells to make proteins, which perform the functions of the virus and continue to replicate themselves.
“Every time a virus replicates itself, they tend to be kind of sloppy at replicating their genetic material. Kind of like really quickly typing out an email without checking your spelling. So, pretty much every time, there’s a mutation or two that gets introduced,” he said. “So if you compare a sample that was collected in January to one that was collected from a different patient a month later, there are now probably two beads that are a different color within the genome.”
By identifying mutations in the genome, the researchers were able to connect different viral strains to outbreaks all over the world.
“We put (the genome samples) into a big analysis and make these family trees that show which variants of the virus are closely related to other strains,” Worobey said. “We found a way to integrate not just the genetic information on the family tree, but also epidemiological information and travel records, which allowed us to do a better job of reconstructing the true history of how these viruses moved.”
In the study, the scientists ran computer programs that simulated the evolution of the virus and showed exactly how it mutated, and where, since the pandemic began.
“By re-running the introduction of SARS-CoV-2 into the U.S. and Europe through simulations, we showed that it was very unlikely that the first documented viral introductions into these locales led to productive transmission clusters,” said study co-author Joel Wertheim of the University of California at San Diego. “Molecular epidemiological analyses are incredibly powerful for revealing transmissions patterns of SARS-CoV-2.”
A blueprint for the future
Worobey and his collaborators agreed that the most important part of this study was the evidence that early, aggressive interventions work to contain viral outbreaks.
“Our research shows that when you do early intervention and detection well, it can have a massive impact, both on preventing pandemics and controlling them once they progress,” Worobey said. “While the epidemic eventually slipped through, there were early victories that show us the way forward: Comprehensive testing and case identification are powerful weapons.”
The team also agreed that the measures taken in those initial cases, which included testing, contact tracing, isolation and cooperation from patients, “should serve as a blueprint for future responses to emerging diseases that have the potential to escalate into worldwide pandemics.”
CDC director: Mask may protect better than Covid-19 vaccine